Prokaryotes and Eukaryotes Revealed: The Unexpected Overlap in the Tree of Life
Prokaryotes and Eukaryotes Revealed: The Unexpected Overlap in the Tree of Life
Highlighting a surprising twist in biological evolution, reveals how long-held distinctions between the simplest microbes and complex cells blur more than once thought. Far from existing in biological isolation, prokaryotes and eukaryotes share a more intricate, interwoven heritage than taxonomy alone suggests—reshaping our understanding of life’s shared ancestry. New genomic and evolutionary studies expose shared genetic pathways, ancient horizontal gene transfers, and unexpected evolutionary back-and-forth, challenging the notion that eukaryotic cells were a sudden leap from prokaryotic origins.
As science peels back layers, the tree of life emerges not as a rigid ladder but as a dynamic, interconnected web.
The Tight Genetic Tether: Shared Genes Challenge Origin Myths
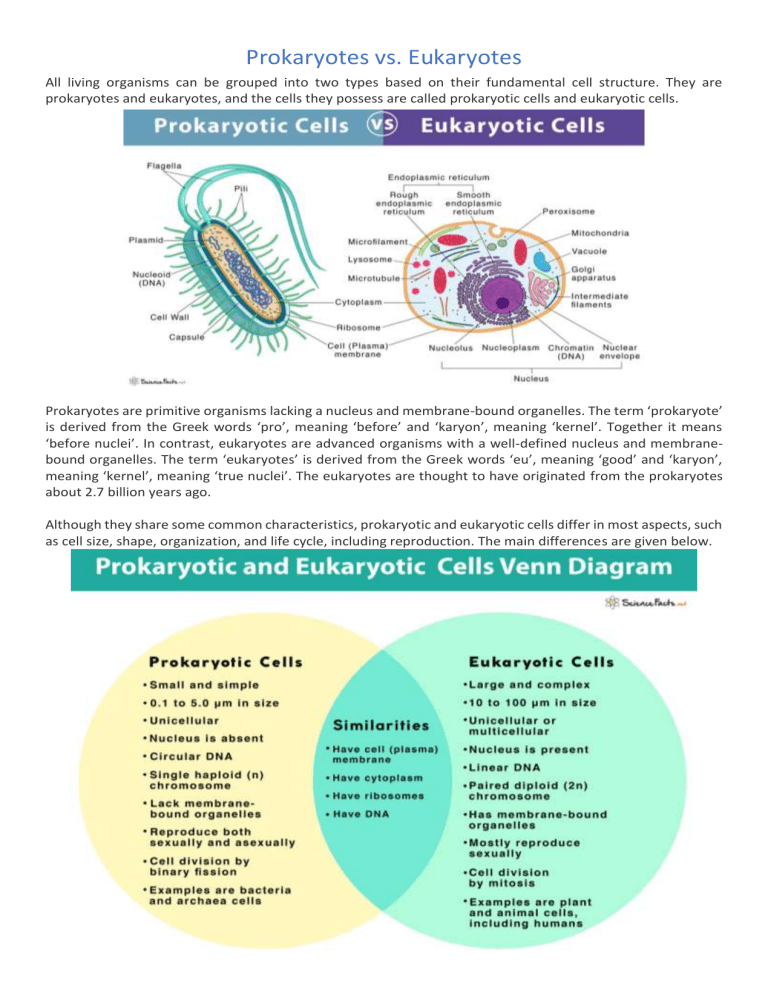
Textbooks long taught that eukaryotes evolved from prokaryotic ancestors via a single, transformative event—likely the endosymbiotic engulfment of bacteria by a host cell. Yet recent evidence reveals this origin was neither isolated nor one-directional. Prokaryotes and eukaryotes possess far more genetic overlap than once assumed: - Core metabolic genes, such as those involved in respiration and energy production, show shared ancestry with both groups.- Horizontal gene transfer (HGT) between archaea and bacteria appears to have seeded early eukaryotic lineages with essential tools for nuclear organization and cellular complexity. - “We’re seeing evidence that eukaryotic innovation wasn’t a sudden jump, but a borrowing and refinement process,” notes microbiologist Dr. Lila Chen of the Max Planck Institute for Microbiology.
“Prokaryotes contributed foundational genes that allowed complex cells to evolve far more rapidly than previously believed.” This genetic intermingling undermines the classic dichotomy, suggesting eukaryogenesis was empowering adaptation through a shared toolkit honed across亿亿 billion microbial years.
The Role of Endosymbiosis Reimagined
The endosymbiotic theory remains central—mitochondria and chloroplasts originated from grassing small prokaryotes inside ancestral cells—but current data add nuance. Instead of a one-time event, multiple waves of endosymbiosis likely contributed to eukaryotic complexity.- Introns and regulatory sequences common in eukaryotes have prokaryotic origins, indicating gene capture long after initial engulfment. - Some eukaryotic lineages still host dynamic prokaryotic symbionts, proving ancient partnerships remain biologically active. - Phylogenomic trees now show eukaryotes weren’t a single fork, but a mosaic settlement shaped by repeated genetic exchanges with diverse microbial partners.
This evolving model supports a “networked origin,” where eukaryotes emerged not from purity, but from a prolonged dialogue with prokaryotic diversity.
A key discovery reshaping phylogeny is the presence of eukaryotic-like genes in archaeal and bacterial genomes—evidence of bidirectional gene flow. These lateral transfers blur species boundaries, suggesting the last universal common ancestor (LUCA) was part of a vibrant, interconnected microbial community rather than a lone progenitor.
One landmark study traced archaic eukaryotic gene homologs in deep-branching bacteria, redefining evolutionary relationships and challenging traditional tree branching. “Prokaryotes and eukaryotes weren’t distant cousins separated by millions of years,” explains evolutionary biologist Dr. Amara Patel.
“They co-opted each other’s genes, blurred phylogenetic lines, and built complexity joint venture-style.” Such findings redefine how scientists interpret evolutionary descent—not as linear descent with modification, but as dynamic integration across cellular domains.
Functional Overlap: From Photosynthesis to Defense
Beyond genetics, biochemical and physiological parallels reveal deeper overlap. Eukaryotic photosynthetic organisms rely on chloroplasts derived from cyanobacteria, yet some protists harbor flexible architectures: they gain chloroplasts via secondary endosymbiosis, then lose or recombine them over time.This plasticity mirrors prokaryotic adaptability, not just eukaryotic simplification. - Eukaryotic immune systems share components with bacterial CRISPR-Cas defenses, indicating ancient acquisition. - Even cellular signaling pathways show prokaryotic-eukaryotic chimeric proteins, suggesting shared regulatory blueprints.
These overlapping functions reveal adaptation strategies evolved independently but accessed through similar molecular foundations—evidence life built complexity not through isolation, but through integration.
Rethinking Evolution: A Web, Not a Ladder
The classic “tree of life” metaphor evokes branching descent, but modern data demand a shift—toward a networked tree reflecting lateral exchange and shared innovation. - Genomic analyses increasingly show hybrids of prokaryotic and eukaryotic traits in ancient lineages.- Fossils and molecular clocks align with a protracted, collaborative evolutionary process, not a sudden leap. - The distinction between prokaryote and eukaryote emerges less as a taxonomic boundary than an ecological continuum shaped by gene sharing, environmental adaptation, and symbiosis. This reimagined lineage underscores life’s shared resilience and creativity—where even the smallest microbes helped construct the blueprint for all complex cells.
Far from being distinct and isolated domains, prokaryotes and eukaryotes reveal a profound, ongoing biological dialogue written into every genome. Their overlapping legacy—woven through genes, metabolism, and symbiosis—demands a revised evolutionary narrative: one where life’s diversity springs not from separation, but from connection.

Related Post

Jon Cryer’s Wife: The Quiet Partnership Behind the Laughter

BMW 330i M Sport 2020: The Telematic Marvel with Proven Performance — Used Specs, Price & Real-World Testimony

Queenzflip Age: Redefining Digital Identity Through Playful Innovation

Terra Luna Classic: What’s the Future Hold? Decoding the Path for a Classic Token Reborn

